Did you know that Jodie Foster gave Penn's commencement address in 2006? And did you know that she raps? Take a look:
Oh man, makes me embarrassed just watching it...
Friday, December 27, 2013
Saturday, December 21, 2013
The over-mathification of Wikipedia
Wikipedia is an amazing resource. Truly amazing. Aside from personal interests, I think I use it at least several times a day for work-related science information. For biology, it's proven to be an invaluable resource for just getting a quick overview of a particular gene or disorder or other biological topic. It could also be a great resource for more theoretical stuff as well, and sometimes it is. For instance, I teach a fluid mechanics class, and one of the main confusions for students is the difference between streamlines, streaklines and pathlines. The Wikipedia page on this topic is just great, and has an animated GIF that gets the point across way better than I can on the chalkboard.
But for many of the theoretical topics, there's a big problem: somewhere along the line, it's clear that some mathematicians got involved. The issue is that they've inserted all sorts of mathematical technicalities into otherwise relatively simple (and useful) mathematical techniques that make the article essentially unreadable and useless. Take the Wikipedia entry on the Laplace transform. The Laplace transform is super useful in solving differential equations, basically because it turns derivatives into multiplication, thereby turning the differential equation into an algebraic equation. Very handy. But good luck getting that out of the Wikipedia article! Instead of starting with a simple application to show how people actually might use the Laplace transform in practice, the Wikipedia article begins with a lengthy and overly mathematical formal definition, with statements like:
Overall, look at the organization of the page. It is just so... mathy! It starts with a formal definition (devoid of any real practical motivation), proceeds to some details about convergence, then a bunch of properties and theorems, then tables of transforms, and then, THEN, finally, some examples. The only part of this that has a bit of motivation for a person who doesn't already know about why the Laplace transform is useful is at the beginning of the "Properties and theorems" section:
But for many of the theoretical topics, there's a big problem: somewhere along the line, it's clear that some mathematicians got involved. The issue is that they've inserted all sorts of mathematical technicalities into otherwise relatively simple (and useful) mathematical techniques that make the article essentially unreadable and useless. Take the Wikipedia entry on the Laplace transform. The Laplace transform is super useful in solving differential equations, basically because it turns derivatives into multiplication, thereby turning the differential equation into an algebraic equation. Very handy. But good luck getting that out of the Wikipedia article! Instead of starting with a simple application to show how people actually might use the Laplace transform in practice, the Wikipedia article begins with a lengthy and overly mathematical formal definition, with statements like:
One can define the Laplace transform of a finite Borel measure μ by the Lebesgue integralWhile these conditions may be interesting to mathematicians, I don't think it is of any interest for the vast majority of people who use the Laplace transform. Then it gets into discussion of the region of convergence, which again begins with:
If f is a locally integrable function (or more generally a Borel measure locally of bounded variation), then the Laplace transform F(s) of f converges provided that the limit...Perhaps you care about the Borel measure locally of bounded variation, but I'm guessing that most people who are interested in the Laplace transform haven't taken courses in point-set topology.
Overall, look at the organization of the page. It is just so... mathy! It starts with a formal definition (devoid of any real practical motivation), proceeds to some details about convergence, then a bunch of properties and theorems, then tables of transforms, and then, THEN, finally, some examples. The only part of this that has a bit of motivation for a person who doesn't already know about why the Laplace transform is useful is at the beginning of the "Properties and theorems" section:
The Laplace transform has a number of properties that make it useful for analyzing linear dynamical systems. The most significant advantage is that differentiation and integrationbecome multiplication and division, respectively, by s (similarly to logarithms changing multiplication of numbers to addition of their logarithms). Because of this property, the Laplace variable s is also known as operator variable in the L domain: either derivative operator or (for s−1) integration operator. The transform turns integral equations and differential equations to polynomial equations, which are much easier to solve. Once solved, use of the inverse Laplace transform reverts to the time domain.
Why is this not at the very beginning? The answer is that it was! Look at this version from 2005. Much better! Still not perfect, since it doesn't have an example, but at least it more clearly gets the main point across about why you would use the Laplace transform. To me, it's clear that at some point, mathematicians got involved and wanted to make the page "right", the same way that mathematicians make the delta function "right" with all kinds of stuff about distributions, in the process completely obscuring the basic point of the delta function for people who really want to use it practically (and many other such mathy topics have been similarly "rightified"). Just to be clear, I'm saying this as a person who has a Ph.D. in math and who loves math, and I think it's great that people far smarter than I have spent time to make sure that one can rigorously define these things. And it IS important, even in some practical contexts. But it's not good for exposition to a more general audience on a Wikipedia page.
Oh, and here's the original Wikipedia page for the Laplace transform. All else aside, we've come a long way, baby!
Oh, and here's the original Wikipedia page for the Laplace transform. All else aside, we've come a long way, baby!
Wednesday, December 18, 2013
Some mathematical principles
I was just thinking about the old days when I used to do math (although not particularly well), and I remember thinking that there are some principles in math. These are general "trueisms" in math that, unlike theorems, are neither provable nor even always true. Actually, I guess then they're "trueishisms". Anyway, here are a couple I know of:
- Conservation of proof energy. I think I heard of this from Robin Hartshorne when I was taking his graduate algebra course. The idea is that if you're trying to prove something hard, and there's a lemma that makes the result easy, then proving that lemma is going to be hard.
- In real analysis, if something seems intuitively true, there's probably a counterexample. For example, the existence of an unmeasurable set. (2 and 3 courtesy of Fang-Hua Lin during his complex variables course, if I remember right.)
- In complex analysis, if something seems too amazing to be true, it probably is true. For example, everything about holomorphic functions.
- In numerical analysis, if you are estimating complexity or error bounds, log(n) for n large is 7 (courtesy of Mike Shelley, I think).
Saturday, December 14, 2013
Lab safety quizzes
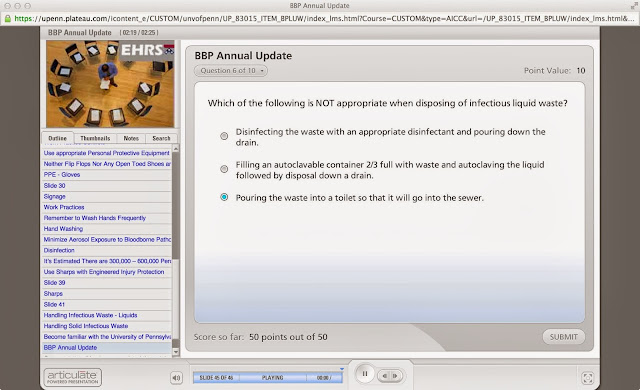
So another year has passed, and I again had to take the online lab safety training, along with the requisite quiz at the end to “test my knowledge of lab safety”. Dunno about you, but I find these quizzes to be completely meaningless: either the questions are ridiculously simple, or they are ridiculously hard, testing some arcane (and ultimately useless) detail of lab safety. Here is an example of the former:
Now you could argue that by having to answer this question, it forces you to read the other two answers and thus learn their content, i.e., “oh, my two legitimate options are autoclaving and disinfecting followed by pouring down the drain.” But then take a look at this example:
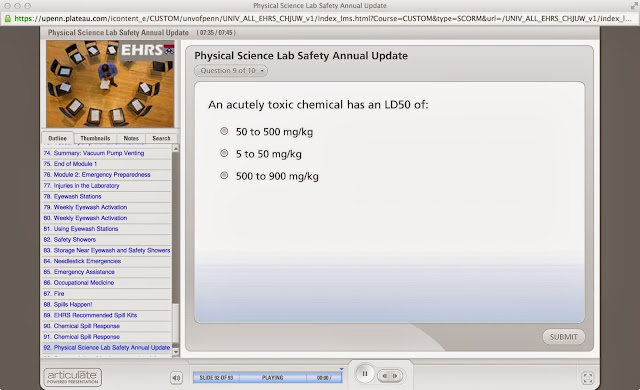
Answers 1, 2, 3 are all obviously wrong (although perhaps its teaching you something about it), but then the one that’s actually correct (80-120 feet per minute face velocity) is actually the one thing I didn’t know at all! It’s like the inverse of the above.
Then you get the ones that are impossible, with arcane answers. I actually encountered a lot more of them when I was at MIT, like the one that asked something like:
EPA regulations dictate that under 40 gallons of oil may be stored in a secondary area for:
1. 3 days
2. 1 week
3. 2 weeks
4. 1 month
Answers 1, 2, 3 are all obviously wrong (although perhaps its teaching you something about it), but then the one that’s actually correct (80-120 feet per minute face velocity) is actually the one thing I didn’t know at all! It’s like the inverse of the above.
Then you get the ones that are impossible, with arcane answers. I actually encountered a lot more of them when I was at MIT, like the one that asked something like:
EPA regulations dictate that under 40 gallons of oil may be stored in a secondary area for:
1. 3 days
2. 1 week
3. 2 weeks
4. 1 month
I think the answer was 3 or something like that. Talk about weird! Here are some similar ones I’ve seen at Penn:
I had absolutely no idea about the lecture bottle explosion question, and the other two I just made educated guesses.
Anyway, I suppose you can say that someone can learn something (albeit very little) from these quizzes, but I certainly don’t think they serve any evaluative purpose. If only they could just teach some basic common sense...
I had absolutely no idea about the lecture bottle explosion question, and the other two I just made educated guesses.
Anyway, I suppose you can say that someone can learn something (albeit very little) from these quizzes, but I certainly don’t think they serve any evaluative purpose. If only they could just teach some basic common sense...
Getting honest feedback
On this blog, I've repeatedly argued that peer review is a net waste of time, basically because it doesn't enforce much quality control, it results in long publication times, and reviewers have seemingly gone mad these days with additional experiments. I say "net waste of time", though, because there are some undeniable benefits. Almost any paper will usually benefit from a well-meaning expert looking through the paper, remarking on which explanations are unclear, which claims are oversold (or undersold), and what further data would be nice to include if you have it.
So if we were to eliminate peer review, where would we get this sort of feedback? Well, ideally, through peer review–that is, actually sending the paper (informally) to your peers and asking for their comments. The problem, of course, is that this is one sort of response:
The problem, as I've documented before, is that nobody has time to read. But I'm lucky to have some good friends who will actually take the time to read a paper and give detailed feedback. For instance, Hyun Youk (postdoc at UCSF) read over a paper Gautham's about to submit and gave us quite detailed and extremely helpful feedback that really strengthened and clarified the paper, like the best peer review ever. I have no idea how to systematize this (I guess our current peer review system is basically that, but anonymous), but it's got me dreaming of a fantasy world where our friends read our papers and love them and make them better and then we post on ArXiV and all get HHMI funding. Sigh. Now to submit this paper and get ready for a war of attrition with the official "peer viewers"...
So if we were to eliminate peer review, where would we get this sort of feedback? Well, ideally, through peer review–that is, actually sending the paper (informally) to your peers and asking for their comments. The problem, of course, is that this is one sort of response:
Hey Arjun,
Nice paper! Very exciting result! My suggestion would be to extend out the discussion a bit and cite these few papers (some papers). Good luck!
Science Friend
The problem, as I've documented before, is that nobody has time to read. But I'm lucky to have some good friends who will actually take the time to read a paper and give detailed feedback. For instance, Hyun Youk (postdoc at UCSF) read over a paper Gautham's about to submit and gave us quite detailed and extremely helpful feedback that really strengthened and clarified the paper, like the best peer review ever. I have no idea how to systematize this (I guess our current peer review system is basically that, but anonymous), but it's got me dreaming of a fantasy world where our friends read our papers and love them and make them better and then we post on ArXiV and all get HHMI funding. Sigh. Now to submit this paper and get ready for a war of attrition with the official "peer viewers"...
Wednesday, December 4, 2013
[Wormpaper outtakes and bloopers] elt-2 RNA level dynamics after heat shock in wild-type and HS::end-1 strains
- Gautham
Second in a series of outtakes and bloopers related to our paper on the relationship of gene expression dynamics and cell division times in the early C. elegans embryo.
In our paper we perturbed cell divisions using mutants and asked if gene expression would track with those cell divisions. Conversely, it would have been great to modify the levels of transcriptional activators and see if gene expression could start before the cell divisions that normally precede them.
Strains with end-1 under the control of a heat shock promoter are available. end-1 is a well-known activator of elt-2. So we went ahead.
Methods: We obtained strain RJ663bc3 from Joel Rothman's group with the kind help of Yewubdar Argaw. We isolated embryos from a synchronized batch culture and aliquoted them at 25C. We heat shocked each aliquot at a different time, for 5 min. at 34C, and returned them to 25C. These brief but accurate heat-shocks were accomplished by centrifuging the embryos and resuspending them in M9 at the appropriate temperature. All samples were fixed simultaneously. The different aliquots spent between 0 and 40 min. at 25C after heat shock and before fixation.
Results: We did RNA-FISH on all the samples for end-1, end-3, and elt-2. We analyzed the data for each aliquot and obtained the following very interesting-looking result for elt-2 expression after end-1 over-expression by heat-shock:
Each panel shows embryos that were heat shocked within a particular window of their development as indicated in the grey shaded region. The red dots are the elt-2 expression levels for these heat-shocked embryos after they've been returned to 25C. The blue dots are the expression trajectory in wild-type without heat-shock for reference and the dashed vertical lines are E lineage divisions. It was super interesting to see elt-2 starting before the 2E divisions, and that it didn't appear to be sensitive if the heat shock came too early.
Blooper: This result is not in the paper because we see qualitatively similar behavior if we heat-shock wild-type worms, with no end-1 overexpressing transgene. Todd Lamitina alerted me to the critical need of doing that control experiment, which in hindsight I should have done first.
Perspective: What was much worse is that I had reason to know that the experiment would be a failure if I had paused to think about all I'd done. Well before doing this experiment I had looked at wild-type embryos that had been left at 30C (not a viable growth temperature for C. elegans) for an hour and found precocious elt-2 expression in these embryos. I disregarded that result as due to any number of things that could go wrong when you try to grow worms under lethal conditions. Without this confounding problem, the overexpression experiment would have been a valuable addition to the paper no matter what the outcome.
Why we were even looking at gene expression trajectories of wild-type worms under non-viable temperatures is a separate story, and another example of Perhaps there will be something interesting.
Second in a series of outtakes and bloopers related to our paper on the relationship of gene expression dynamics and cell division times in the early C. elegans embryo.
elt-2 RNA level dynamics after heat shock in wild-type and HS::end-1 strains
This one is a blooper.In our paper we perturbed cell divisions using mutants and asked if gene expression would track with those cell divisions. Conversely, it would have been great to modify the levels of transcriptional activators and see if gene expression could start before the cell divisions that normally precede them.
Strains with end-1 under the control of a heat shock promoter are available. end-1 is a well-known activator of elt-2. So we went ahead.
Methods: We obtained strain RJ663bc3 from Joel Rothman's group with the kind help of Yewubdar Argaw. We isolated embryos from a synchronized batch culture and aliquoted them at 25C. We heat shocked each aliquot at a different time, for 5 min. at 34C, and returned them to 25C. These brief but accurate heat-shocks were accomplished by centrifuging the embryos and resuspending them in M9 at the appropriate temperature. All samples were fixed simultaneously. The different aliquots spent between 0 and 40 min. at 25C after heat shock and before fixation.
Results: We did RNA-FISH on all the samples for end-1, end-3, and elt-2. We analyzed the data for each aliquot and obtained the following very interesting-looking result for elt-2 expression after end-1 over-expression by heat-shock:
Each panel shows embryos that were heat shocked within a particular window of their development as indicated in the grey shaded region. The red dots are the elt-2 expression levels for these heat-shocked embryos after they've been returned to 25C. The blue dots are the expression trajectory in wild-type without heat-shock for reference and the dashed vertical lines are E lineage divisions. It was super interesting to see elt-2 starting before the 2E divisions, and that it didn't appear to be sensitive if the heat shock came too early.
Blooper: This result is not in the paper because we see qualitatively similar behavior if we heat-shock wild-type worms, with no end-1 overexpressing transgene. Todd Lamitina alerted me to the critical need of doing that control experiment, which in hindsight I should have done first.
Perspective: What was much worse is that I had reason to know that the experiment would be a failure if I had paused to think about all I'd done. Well before doing this experiment I had looked at wild-type embryos that had been left at 30C (not a viable growth temperature for C. elegans) for an hour and found precocious elt-2 expression in these embryos. I disregarded that result as due to any number of things that could go wrong when you try to grow worms under lethal conditions. Without this confounding problem, the overexpression experiment would have been a valuable addition to the paper no matter what the outcome.
Why we were even looking at gene expression trajectories of wild-type worms under non-viable temperatures is a separate story, and another example of Perhaps there will be something interesting.
[Wormpaper outtakes and bloopers] Expression dynamics in C. elegans mes-2 mutant embryos
- Gautham
Very recently I got into a discussion in which I was being super negative about our tendency as scientists to seek to do "one more experiment," and the feeling that it is possible to increase the impact of work by doing more of it. Frankly, if you look at papers in "top tier" journals we see papers that appear to be a collection of somewhat unrelated results bundled up into a massive effort. So it definitely feels like a necessary evil to survive in academia, and even in our lab you'll hear folks complaining about how "thin" such-and-such paper is. The painful part is that the drive to avoid a "thin" paper often ends up in inconclusive or negative results which are never published.
This morning I woke up thinking that instead of being so damn negative (which is depressing for morale of all involved), we could do something positive and actually put some of that stuff up on the web, something Arjun's been encouraging since he set up the blog but we haven't really followed up with in the group. That way the work wasn't for nothing and even though its not in a journal, maybe google will find it when someone makes a search.
This is the first of a few short posts on experiments that we did in relation to our paper on the relationship between cell division times and gene expression onsets in early development of C. elegans embryos. The core of that paper was completed rather quickly, but we spent quite some time trying to add stuff to it. Here is some of the stuff that didn't make the cut.
Discussion: Seeing no effect was disappointing, but it doesn't necessarily contradict the Yuzyuk report, since most of its statements on expression level effects deal with 8E or later stages in development. That is right about where our window of interest ended.
Perspective: I did these experiments on the common rationale that Perhaps there would be something interesting. That is a good reason to do an experiment. In fact, that is how all experiments get started.
However, it was a bad reason to withhold developing the manuscript for the actual paper, because this experiment, no matter what the outcome would have been, does not have all that much to do with the question of how cell divisions and expression timing are coordinated. So that should have continued as a parallel rather than an in-series effort. What is very telling is that this result is not in the paper because it was a negative result, but we would have probably found a way to put it in if it was positive. Very few experiments of that sort are related in an honest way to the paper that they are being bundled with. Its just fluff.
But we waited because of that feeling that bundling cool stuff together makes for a more compelling and publishable paper. And because, currently, there is no home for "thin" results, like this little blurb on mes-2.
Very recently I got into a discussion in which I was being super negative about our tendency as scientists to seek to do "one more experiment," and the feeling that it is possible to increase the impact of work by doing more of it. Frankly, if you look at papers in "top tier" journals we see papers that appear to be a collection of somewhat unrelated results bundled up into a massive effort. So it definitely feels like a necessary evil to survive in academia, and even in our lab you'll hear folks complaining about how "thin" such-and-such paper is. The painful part is that the drive to avoid a "thin" paper often ends up in inconclusive or negative results which are never published.
This morning I woke up thinking that instead of being so damn negative (which is depressing for morale of all involved), we could do something positive and actually put some of that stuff up on the web, something Arjun's been encouraging since he set up the blog but we haven't really followed up with in the group. That way the work wasn't for nothing and even though its not in a journal, maybe google will find it when someone makes a search.
This is the first of a few short posts on experiments that we did in relation to our paper on the relationship between cell division times and gene expression onsets in early development of C. elegans embryos. The core of that paper was completed rather quickly, but we spent quite some time trying to add stuff to it. Here is some of the stuff that didn't make the cut.
Expression dynamics in C. elegans mes-2 mutant embryos
In the midst of working on the project that resulted in the paper I went to a talk by Prof. Susan Mango describing their work on mes-2 mutants (Yuzyuk et al. Dev Cell 2009). MES-2 is a component of Polycomb and its mutation was reported to change the window of developmental plasticity in the embryo. I thought it would be interesting to measure the dynamics of the genes to be featured in our wormpaper in this mutant background, since the paper was all about timing of expression of lineage-specification genes. Perhaps there would be something interesting.
Methods: mes-2(bn11) was the mutant studied by Yuzyuk et al. We got it in strain SS186 from the C. Elegans Genetics Center. mes-2 mutants have a very peculiar phenotype: Progeny of mes-2 homozygous mothers are sterile. We switched the balancer with a GFP balancer by mating with strain MT20110, constructed by Erik Andersen, a friend and collaborator of Arjun's from his postdoc time. I wanted to avoid using a fluorescent worm-sorter, so I ended up manually removing all GFP(+) worms from a small synchronized plate. The remaining worms are either fertile mes-2 first-generation homozygotes or their sterile progeny. Very carefully, we ran those ~1000 worms through a micro-scale version of our worm embryo preparation (which usually works with >10x the amount of worms). We then conducted RNA-FISH as usual.
Results: In the developmental window we were interested in there were no changes at all in the RNA level dynamics of genes we tested compared to wild type (N2 strain). Below is a figure for our favorite genes: end-1, end-3, and elt-2.
Similarly, in our hands, the expression dynamics of elt-7, hlh-1, and elt-1, were nearly identical to N2.
Discussion: Seeing no effect was disappointing, but it doesn't necessarily contradict the Yuzyuk report, since most of its statements on expression level effects deal with 8E or later stages in development. That is right about where our window of interest ended.
Perspective: I did these experiments on the common rationale that Perhaps there would be something interesting. That is a good reason to do an experiment. In fact, that is how all experiments get started.
However, it was a bad reason to withhold developing the manuscript for the actual paper, because this experiment, no matter what the outcome would have been, does not have all that much to do with the question of how cell divisions and expression timing are coordinated. So that should have continued as a parallel rather than an in-series effort. What is very telling is that this result is not in the paper because it was a negative result, but we would have probably found a way to put it in if it was positive. Very few experiments of that sort are related in an honest way to the paper that they are being bundled with. Its just fluff.
But we waited because of that feeling that bundling cool stuff together makes for a more compelling and publishable paper. And because, currently, there is no home for "thin" results, like this little blurb on mes-2.
Code reuse and plagiarism
Gautham's been doing a bunch of refactoring of our existing codebase for spot counting, in particular using a lot of object oriented design strategies. One of the primary goals is code reuse, which is commonly accepted as A Good Thing in programming. On the other hand, a student in lab has been writing a grant proposal on a topic that is very similar to another grant proposal we have submitted (and yes, we checked with the funding agency, it's okay in this context). But of course, my student felt compelled to have completely new language so that it wasn't "plagiarism", which is A Bad Thing. Which got me wondering: why are we so concerned about plagiarism in this context? Why is reuse of language in one context a worthy goal in and of itself, and complete blasphemy in another? Here's another example: I was at a thesis defense in which the candidate was strong second author on a paper and had included some of the figure legends from the published paper in the figures in her thesis. One of the committee members complained about this, saying that it was important to "write this in one's own words". I agree that it may be a good exercise, but I'm not convinced that using the figure legends is per se A Bad Thing. There just aren't all that many clear and concise ways of writing the same thing.
Maybe it's time to rethink the plagiarism taboo. Just like you can use someone else's computer code (with appropriate attribution), why not be able to use someone else's language? If someone wrote something really well, what's the point in rewriting it–probably less well–just for the sake of rewriting it? Would anyone rewrite a super efficient algorithm just for the sake of saying "I wrote it myself"? All that you need is a good mechanism for attribution, like quotes and links and stuff, you know, like all that stuff we already use in writing. In fact, I would argue that attribution is far more transparent in writing than in computer code, because typically only the programmer sees the code, not the end user. If I run a program, I typically am not looking at the source code, so I don't really know who did what, even if the README file gives credit to other code sources.
One might object to copy-and-paste writing becoming a mish-mash of ill-fitting pieces. First off, this is just bad writing, just as code that just jams together different pieces can be a mess, and will typically require one to code stuff to make things flow together well. But in a world where writing demands are growing daily (certainly the case for me), maybe it's time to consider text-reuse as a good practice, or at least not necessarily a bad one.
Saturday, November 16, 2013
What is the best text editor?
Ah, why a post on this, when there are probably already something like 17,000 posts being written on the topic at any given time? Well, it’s a topic we often debate in lab, usually as a cool down from some protracted Python vs. MATLAB vs. Perl vs. R argument. Gautham likes vi, as do many programming junkies. I used to use Emacs for a while, then migrated to a little known text editor called Smultron, and now I use some combination of the IDEs incorporated in MATLAB and R and TextWrangler for everything else. And you know what? It works! I can actually edit text this way! I can save the changes, and the run the program! Amazing. What’s even more amazing is that virtually every text editor does this. The conclusion I came to is that it just doesn’t much matter how many fancy key codes your editor can handle (looking at you, Emacs!), nor how complex or simple the interface is (to a point, and once you get used to it), because unless you’re working with punchcards, your programming is much more likely to be limited by your mind than your choice of text editing program. I used to think more about text editors back when I was in graduate school, and I remember one day stopping by the office of Boyce Griffith, who is one of my academic “brothers” (i.e., we had the same advisor). Now, Boyce is one of the most incredible scientific programmers I’ve ever met, having written literally millions of lines of hard core C and C++ to implement very complex simulations of fluids, including these amazing simulations of the human heart. I saw Boyce was programming away using some fairly primitive text editor, and I was like “Man, you don’t use Emacs or something?” Boyce said “No”, and I said “Why not?” and he said “Whatever, it doesn’t matter.” Amen. In the end, I think the best text editor is, you know, one that edits text.
Tuesday, November 12, 2013
some thoughts related to "The Frustrated Gene: Origins of Eukaryotic Gene Expression"
- Gautham
Chris forwarded this interesting essay to the group:
The Frustrated Gene: Origins of Eukaryotic Gene Expressionby Hiten D. Madhani.
The arguments are not airtight, but how could they be anyway? A friend of mine has said that these efforts are necessarily a connect-the-dots exercise with the dots spread very far apart. It is not at all easy to figure out how these ancient devices came about. Nevertheless, the presented idea that the complexity of gene regulation was created to temporarily ward off selfish DNA makes more sense than it having arisen primarily to actually enable complexity. I think that the perspective still leaves open the possibility that we've kept all those adaptations because they do enable complexity, but I am nowhere near qualified to judge that for sure.
What is clear is that looking at every aspect of gene regulation in eukaryotes and thinking that it has to create a net benefit to the organism at the present, rather than maybe resulting from a prior bottleneck, a prior arms race with a parasite, or even the defeat at the hands of a parasite, may be a mistake. It is terrifying how natural selection is rather helpless in preventing the spread of even damaging transposons in a sexual population. Biology is supposed to make sense in light of evolution, but it may be that no organism may make sense if we require it to be in all aspects evolutionary optimal.
I recall one of Marc Kirschner's book saying that it was at the time an ongoing embarrassment to evolutionary biologists that they don't have a solid explanation for why sexual reproduction is advantageous. They give us the simple story that it enabled complexity and evolvability, but they themselves are not convinced that the numbers work out, or at least they weren't a decade or two ago. It is hard for us to even make a case for sexual reproduction, and yet some of us are losing sleep trying to explain things like apparent gene redundancy.
It is possible that us larger eukaryotes at least operate in too big a variety of circumstances, and perhaps we compete more on the basis of behavior than on the basis of physiology. Perhaps our environment changes too rapidly, and we reproduce too slowly, to allow us to reach our biochemical evolutionary pinnacle. I've been watching "The Life of Mammals" series from BBC. It seems far easier to explain the extraordinary diversity in behavior and shapes of animals on evolutionary grounds, than it is to explain even basic things like their wildly different DNA content.
Chris forwarded this interesting essay to the group:
The Frustrated Gene: Origins of Eukaryotic Gene Expressionby Hiten D. Madhani.
The arguments are not airtight, but how could they be anyway? A friend of mine has said that these efforts are necessarily a connect-the-dots exercise with the dots spread very far apart. It is not at all easy to figure out how these ancient devices came about. Nevertheless, the presented idea that the complexity of gene regulation was created to temporarily ward off selfish DNA makes more sense than it having arisen primarily to actually enable complexity. I think that the perspective still leaves open the possibility that we've kept all those adaptations because they do enable complexity, but I am nowhere near qualified to judge that for sure.
What is clear is that looking at every aspect of gene regulation in eukaryotes and thinking that it has to create a net benefit to the organism at the present, rather than maybe resulting from a prior bottleneck, a prior arms race with a parasite, or even the defeat at the hands of a parasite, may be a mistake. It is terrifying how natural selection is rather helpless in preventing the spread of even damaging transposons in a sexual population. Biology is supposed to make sense in light of evolution, but it may be that no organism may make sense if we require it to be in all aspects evolutionary optimal.
I recall one of Marc Kirschner's book saying that it was at the time an ongoing embarrassment to evolutionary biologists that they don't have a solid explanation for why sexual reproduction is advantageous. They give us the simple story that it enabled complexity and evolvability, but they themselves are not convinced that the numbers work out, or at least they weren't a decade or two ago. It is hard for us to even make a case for sexual reproduction, and yet some of us are losing sleep trying to explain things like apparent gene redundancy.
It is possible that us larger eukaryotes at least operate in too big a variety of circumstances, and perhaps we compete more on the basis of behavior than on the basis of physiology. Perhaps our environment changes too rapidly, and we reproduce too slowly, to allow us to reach our biochemical evolutionary pinnacle. I've been watching "The Life of Mammals" series from BBC. It seems far easier to explain the extraordinary diversity in behavior and shapes of animals on evolutionary grounds, than it is to explain even basic things like their wildly different DNA content.
Thursday, October 31, 2013
Sequencing is the new pcr
- Gautham
I was wondering if the domination of high-throughput sequencing in genome science is an anomaly of the times, so I decided to look at the table of contents of old Genome Research issues. What was Genome Research about before the advent of all this sequencing?
http://genome.cshlp.org/content/2/1.toc
http://genome.cshlp.org/content/1/4.toc
Take a look! It was even more dominated by PCR then than it is dominated by HT sequencing now.
I was wondering if the domination of high-throughput sequencing in genome science is an anomaly of the times, so I decided to look at the table of contents of old Genome Research issues. What was Genome Research about before the advent of all this sequencing?
http://genome.cshlp.org/content/2/1.toc
http://genome.cshlp.org/content/1/4.toc
Take a look! It was even more dominated by PCR then than it is dominated by HT sequencing now.
Wednesday, October 30, 2013
Better programming practices applied to a scientific plotting problem
- Gautham
It is easy to find "best programming practices" out on the web, but usually there aren't accompanied by an example. This is a real-life example of applying some good practices to turn a bewildering task into something actually pleasant.
One of the hardest aspects of making the worm paper was dealing with the complicated plots that had to be generated. The input data was of exactly two forms:
You see here lineages versus time (bottom left), RNA versus time (bottom center) and versus nuclei (top right), and even a nuclei versus time plot (bottom right). RNA traces for different genes are shown, color-coded, and a specific lineage (the E lineage) is highlighted and the times of its divisions are highlighted in all the plots. Plus there's the shaded alternating grey bars.
My first crack at this thing was for our first submission. It involved very messy and long scripts in MATLAB. Producing plots required a lot of looking back and forth and copying and pasting. Changing which lineage data or which RNA-FISH data set was used to make the plot was a chore. When the reviews came back, and we had to consider many ways to present our data to make our point clear, we needed a more flexible way to make these plots. Here are some principles/tips that helped me:
Centralize access to raw data
This is related to a previous post on keeping together all data of the same type.
LINDATADIR = 'lineagedata/';
RNADATADIR = 'RNAdata/';
lindata = loadWormLineageData( LINDATADIR );
rnadata = loadAllWormRNAData( RNADATADIR );
>> showIndexWormData(rnadata.gut)
1) RT 101003
2) 15C 101011
3) RT 101115
...
49) N2 20C 110219 and 120309 gut
rnadatacell = { ...
getWormData( rnadata.gut, 'N2 20C 110219' ), ...
getWormData( rnadata.gut, 'N2 20C 110219' ), ...
getWormData( rnadata.gut, 'N2 120216 20C' )};
genenamecell = {'end3','end1','elt2'};
dyenamecell = {'alexa','tmr','cy'};
RNArangecell = {[-100 700],[-100 900],[-100 600]};
lineagedata = lindata.Bao_to20C(1);
figure(3);clf;
plotfig1_0114( rnadatacell, genenamecell, ...
dyenamecell, RNArangecell, lineagedata)
It is easy to find "best programming practices" out on the web, but usually there aren't accompanied by an example. This is a real-life example of applying some good practices to turn a bewildering task into something actually pleasant.
One of the hardest aspects of making the worm paper was dealing with the complicated plots that had to be generated. The input data was of exactly two forms:
- RNA-FISH data sets. A sample of worm embryos of some particular strain, grown under particular conditions, were fixed and imaged. For each embryo analyzed you have #nuclei, and the number of RNA-FISH spots in all fluorescent dye channels used in that experiment. Each dye channel corresponds to a different gene, and different data sets can involve different genes.
- Embryo cell lineage data sets. Our collaborators Travis Walton and John Murray took movies of individual embryos from certain strains and conditions and determined the time of each cell division. We also incorporated published lineage data. The names of the cells in the embryo follow a standardized pattern.
Since the C. elegans embryonic lineage is invariant, you can use the data set in 2 to effectively determine a time axis for RNA-FISH, and do things like determine whether expression of a gene starts before or after a certain cell's division (which was actually a key question asked by the paper).
The goal is to make plot layouts like:
 |
| Output of the figure-1 producing code in MATLAB |
My first crack at this thing was for our first submission. It involved very messy and long scripts in MATLAB. Producing plots required a lot of looking back and forth and copying and pasting. Changing which lineage data or which RNA-FISH data set was used to make the plot was a chore. When the reviews came back, and we had to consider many ways to present our data to make our point clear, we needed a more flexible way to make these plots. Here are some principles/tips that helped me:
Centralize access to raw data
This is related to a previous post on keeping together all data of the same type.
RNADATADIR = 'RNAdata/';
rnadata = loadAllWormRNAData( RNADATADIR );
>> rnadata
rnadata =
gut: {1x49 cell}
othergut: {1x8 cell}
muscle: {1x12 cell}
rnadata =
othergut: {1x8 cell}
muscle: {1x12 cell}
>> showIndexWormData(rnadata.gut)
1) RT 101003
2) 15C 101011
3) RT 101115
...
49) N2 20C 110219 and 120309 gut
In Matlab, structs or arrays of structs are the container of choice for arbitrary data. Above I have a cell array of structs for each experiment, and each struct contains a string that has a descriptor of the experiment, which I can show with the showIndexWormData function. Similarly,
>> lindata
lindata =
n2: [1x1 struct]
gg52: [1x3 struct]
gg19: [1x3 struct]
div1: [1x5 struct]
...
lindata =
gg52: [1x3 struct]
gg19: [1x3 struct]
div1: [1x5 struct]
...
In this case lindata is itself a struct, where each field corresponds to a strain or source and contains a struct array of lineage data, which could represent data from different individual embryos.
Good code needs few comments
If you have to write many comments to explain what your code does, rewrite the code. Code that makes its intent clear is good code. The nightmare is not looking back at code and asking "how does this code work?" but rather "what was I trying to do here!?"
Here is the top level script run to make the figure:
%% FIGURE 1 - Overview
% The end-3 and end-1 data is in 110219
% The elt-2 data is in 120216
getWormData( rnadata.gut, 'N2 20C 110219' ), ...
getWormData( rnadata.gut, 'N2 20C 110219' ), ...
getWormData( rnadata.gut, 'N2 120216 20C' )};
dyenamecell = {'alexa','tmr','cy'};
RNArangecell = {[-100 700],[-100 900],[-100 600]};
plotfig1_0114( rnadatacell, genenamecell, ...
dyenamecell, RNArangecell, lineagedata)
set(gcf,'PaperPositionMode','auto')
print('DRAFT_fig1_0114.eps', '-depsc')
If clarity of intent is key, we are prepared to sacrifice all manner of other things, including speed, storage and efficiency. As you can see, the script really packages the variables I might be interested in changing when producing Fig. 1 and sends them off to a function that does the work.
One other good reason to have such self-commented code is that your code and your comments will never diverge due to laziness. If you rely on comments to understand your code once, you will have to be diligent from then on to make sure they are totally sync'ed up any time you come back and modify it.
Use accessor and constructor functions liberally
This is an offshoot of ideas related to the way you write object-oriented programs. Marshall Levesque turned me onto writing "get" and "make" functions.
getWormData( rnadata.gut, 'N2 20C 110219' )
Fetches data by descriptor, rather than by number, filename, or some other obtuse thing.
Here is the portion of the code in plotfig1_0114 that plots the three plots in the bottom center of the figure:
for i=1:length(rnadatacell)
axes(ht(i));
xlim(RNArangecell{i})
addtimeshading_t( getLinDatabycellname('ABa',lindata_adj) , ...
10, brightcolor, darkcolor, 'y_t');
plot_onewormseries_vs_t( rnadatacell{i}, lindata_adj,...
getRNAslopes( genenamecell{i}),...
getWormColors( genenamecell{i}), dyenamecell{i}, 'x_RNA__y_t') ;
addLineageMarks_t( lindata_adj, Linsubsetfunc, 'y_t') ;
axes(ht(i));
xlim(RNArangecell{i})
addtimeshading_t( getLinDatabycellname('ABa',lindata_adj) , ...
10, brightcolor, darkcolor, 'y_t');
plot_onewormseries_vs_t( rnadatacell{i}, lindata_adj,...
getRNAslopes( genenamecell{i}),...
getWormColors( genenamecell{i}), dyenamecell{i}, 'x_RNA__y_t') ;
addLineageMarks_t( lindata_adj, Linsubsetfunc, 'y_t') ;
end
There are three 'get's here: getLinDatabycellname('ABa',lindata_adj) finds the time of cell division of cell ABa based on the lineage data and getWormColors( genenamecell{i}) returns a structure containing all the graphical properties I like to use when plotting that particular gene. This is how a consistent color is applied every time for a given gene (green for elt-2, for example). In my implementation, the table of graphical properties for all genes is hard coded into that function body and all it does is look up the entry for the desired gene. ( getRNAslopes is also a "get" involving a hard-coded table )
Incidentally, the low level functions accept things like the strings 'x_RNA__y_t' or 'y_t' to determine what is plotted on the x and y axes. This was important when we were considering all kinds of variations with this or that plot flipped.
The lineage marks are added using the lineage data and a function that filters only the lineage I wanted to highlight (the E lineage including the EMS division):
addLineageMarks_t( lindata_adj, Linsubsetfunc, 'y_t')
Turns out that the filter itself was made with an accessor/constructor function:
Linsubsetfunc = makeLinsubsetter('EwEMS');
Here is a function that returns a function itself. MATLAB is a bit handicapped for this kind of thing since the only kind of function you can make interactively is an anonymous function one-liner, but it sufficed for my purpose:
function fh = makeLinsubsetter( lineagename )
switch lineagename
case 'AB'
fh = @(cellnames) find( strncmp('AB',cellnames,2) );
case 'E'
fh = @(cellnames) find( strncmp('E',cellnames,1) & ...
~ strcmp('EMS',cellnames) );
...
strncmp('P0',cellnames,2) | strncmp('NA',cellnames,2)));
case 'all'
fh = @(cellnames) 1:length(cellnames);
% Return a function handle to an anonymous function that will filter for
% the specified lineage.
case 'AB'
fh = @(cellnames) find( strncmp('AB',cellnames,2) );
case 'E'
fh = @(cellnames) find( strncmp('E',cellnames,1) & ...
~ strcmp('EMS',cellnames) );
...
case 'EwEMS'
fh = @(cellnames) find( strncmp('E',cellnames,1));
...
case 'notABorP0'
fh = @(cellnames) find( ~ ( strncmp('AB',cellnames,2) | ...strncmp('P0',cellnames,2) | strncmp('NA',cellnames,2)));
case 'all'
fh = @(cellnames) 1:length(cellnames);
Having a function like this was incredibly handy.
Avoid copy-paste code. Wrapper functions are nice.
These two principles are closely related. For example, consider the plotting of RNA vs time (bottom center) and the RNA vs nuclei (center left) in the figure. The function that plots an individual RNA vs time graph is:
function [ linehandles, plothandles ] = plot_onewormseries_vs_t( cdata, ...
lindata, rnaslopes, graphicsspecs, colortoplot, plotmode)
...
...
embryoages = assignAgesFrom_lindata( numcells, ...
numrna, rnaslopes, lindata) ;
linehandles = plotGoodBadWormData( embryoages, numrna, ...
has_clearthreshold, graphicsspecs, flip_rna_t) ;
lindata, rnaslopes, graphicsspecs, colortoplot, plotmode)
...
...
embryoages = assignAgesFrom_lindata( numcells, ...
numrna, rnaslopes, lindata) ;
linehandles = plotGoodBadWormData( embryoages, numrna, ...
has_clearthreshold, graphicsspecs, flip_rna_t) ;
end
On the other hand, the code that plots an individual RNA vs nuclei graph is:
function [linehandles, plothandles] = plot_onewormseries_vs_N( cdata, ...
graphicsspecs, colortoplot, plotmode)
...
...
graphicsspecs, colortoplot, plotmode)
...
...
linehandles = plotGoodBadWormData( numcells, numrna, ...
has_clearthreshold, graphicsspecs, flip_rna_n) ;
has_clearthreshold, graphicsspecs, flip_rna_n) ;
end
Both functions ultimately call plotGoodBadWormData to actually draw the graphs, but the time version passes in embryo ages, which it computes, rather than the number of cells (same as the number of nuclei). This way, plotGoodBadWormData does the work of plotting and applying all the graphics parameters and its code does not have to be duplicated. Meanwhile, the two plot_onewormseries... functions are really just wrappers.
Ultimately, in all my worm code there are only three or four functions that do actual hard "work".
More function, less scripting.
This is a corollary of avoiding code duplication. Think twice before you copy-paste! If you are using a language like R or python that allows you to write arbitrary functions interactively, there is no down-side to functionalizing.
Hope some of these tips help you to not lose your mind, like I did on my first go at this stuff. We scientists are some pretty bad programmers because nobody else usually sees our code. But we have to see it and work with it. Good practices do make you more efficient, and also, happier. I trust you have enough good taste to avoid the extremes.
Tuesday, October 29, 2013
The design police
Design is everywhere these days. It’s got to be the Apple effect: they’ve definitely pushed design into the public consciousness, and now we’re stuck listening to these insufferable design-nerds anytime any website or software is updated anywhere, no matter how inconsequential the change. Perhaps it’s ironic that Apple itself is the target of most of this design-related hate-babble, with literally every aspect of their software being the subject of endless rants about stuff like kerning and “icon balance” and other stuff nobody cares about.
You know the real issue here? Apple already has great design. Design-police people: please turn your attention to the rest of us! We need you over here! Could iOS7 use slightly less transparency here or there? Maybe. But have you looked at the upenn.edu website anytime lately? Especially the functional parts that us members of the university have to use almost daily? Or our fancy new Concur expense reporting system that borders on useless? Trust me, there’s tons to do because practically the rest of the world needs a design overhaul. I wish the world I worked in most of the time was even remotely as well designed as iOS7 (or frankly, iOS1). And perhaps you could train us on figure making while you’re at it?
By the way, I should say that I do actually appreciate good design, despite it all (and I think iOS7 is actually pretty great). One of my favorite movies on the topic is Helvetica, about, you know, the font. Basically a bunch of design guys getting on there talking about how Helvetica is overused and boring and bad. One of my favorite parts of the movie, though, is when they interview these two design guys and one says something like: “Look, just make your text in Helvetica Bold and it will look good.” Amen.
Saturday, October 26, 2013
Product differentiation when there is no difference
(No, I don't mean this kind of product differentiation.)
I was just reading this article in the NYtimes.com about how sales of bottled water will soon surpass that of soda. Of course, this is utterly ridiculous. It's &*#$-ing water! You know, the stuff that flows from your tap almost for free? Whatever, I guess that's an old argument, and is seems that side of rationality has lost that one. But one of the interesting things in the article was about how all these water companies are trying to differentiate themselves to get an edge because the plain old bottled water market is so competitive that there's very little profit left:
I was just reading this article in the NYtimes.com about how sales of bottled water will soon surpass that of soda. Of course, this is utterly ridiculous. It's &*#$-ing water! You know, the stuff that flows from your tap almost for free? Whatever, I guess that's an old argument, and is seems that side of rationality has lost that one. But one of the interesting things in the article was about how all these water companies are trying to differentiate themselves to get an edge because the plain old bottled water market is so competitive that there's very little profit left:
The TalkingRain Beverage Company, a bottled water business that started in the Pacific Northwest, is getting out of the plain water business altogether because the economics are so bad. “The water business has become very commoditized,” said Kevin Klock, its chief executive.(Umm, "very commoditized"?!?! Sorry, gotta say it again: It's &*#$-ing water!)
Which got me thinking about what really is a commodity. Even tap water does taste different in different places, and I suppose some bottles might be easier to open than others. Grains, chemicals, minerals–they can all come in different purities and grades. In fact, even in lab, we buy all sorts of different types of water for different purposes, including, umm, bottled RNase free water (oops!). So whatever the perfect commodity is, it would have to be something digital and thus inherently pure, like some form of information. But it can't just be all information. For instance, if you're selling some kind of information, you're probably selling information that's hard to come by or process or present in some way. So then I'm not sure if that qualifies as a commodity.
Which leaves money. Money is just a number, and it's equally valuable wherever you go or from whoever you get it; there is no different kind of money (there are financial schemes and categories of investments, but they are not money in and of itself). Of course, you might be wondering who would be selling such a "product". Like, who's out there saying "here's an authentic $10 bill, I'll sell it to you for $10.50!" Well, that's exactly what credit cards do, for instance. So how do different cards differentiate themselves? How do you differentiate yourselves when you're just giving people money? Seems like there are two ways they do it. One is psychological manipulation. These include making merchants pay more or less, which is essentially a hidden cost to the consumer, and those insidious schemes that play upon people's "just pay later" mentality. The other differentiator is the various bonus rewards schemes, which rely on other companies' real world products to provide differentiation.
But what surprises me is that they don't really seem to compete on cost. As in, they ALL have astronomical interest rates. Now, for something that is to my mind as commoditized as a commodity could be, that seems quite strange. Apparently, you can get bottled water for as little as 8 cents per bottle in bulk because competition has driven enormous efficiencies in the production of the "product" (i.e., bottle manufacturing). Why is there so little price competition in credit cards? Sure, on interest rates, they have to cover the folks who don't pay their bills, and on charging businesses, they have to cover the infrastructure costs. Still, shouldn't one company be able to ultimately offer much lower rates than another? I think the clearest answer is cartel behavior. And indeed, they have been hit with many such fines for that sort of behavior in the past. I think it's because the only way to make money (and tons of it) in a purely commodity business is to cheat. In that way, you could view the price competition for water as a shining example of the highs and lows of capitalism: competition drives incredible optimization and efficiencies in a product whose very existence is essentially ridiculous.
Definitely don't want to step into any arguments about the pros and cons of capitalism and regulation. But I will offer up this funny anecdote that might provide fodder for both sides of the argument. I was in DC some time ago for a grant panel, and in the room, they had... bottled water, of course. But apparently, they weren't allowed to call it bottled water, most likely because of some well-intentioned rule against bottled water. So instead, they had these bottles specially labeled by the caterer as "refreshing drink" or something like that, which (ironically) probably cost more than just getting regular old bottled water. Sigh. I was in Canada for a conference recently, and they had a bunch of cups and a pitcher. Now that was refreshing.
Definitely don't want to step into any arguments about the pros and cons of capitalism and regulation. But I will offer up this funny anecdote that might provide fodder for both sides of the argument. I was in DC some time ago for a grant panel, and in the room, they had... bottled water, of course. But apparently, they weren't allowed to call it bottled water, most likely because of some well-intentioned rule against bottled water. So instead, they had these bottles specially labeled by the caterer as "refreshing drink" or something like that, which (ironically) probably cost more than just getting regular old bottled water. Sigh. I was in Canada for a conference recently, and they had a bunch of cups and a pitcher. Now that was refreshing.
Thursday, October 10, 2013
Some analogies for the Higgs field
I was just reading about the Nobel Prize in Physics on NYTimes.com, which this year was for the discovery of the Higgs boson, and came across the following analogy for how the Higgs imbues particles with mass:
"According to this model, the universe brims with energy that acts like a cosmic molasses, imbuing the particles that move through it with mass, the way a bill moving through Congress attracts riders and amendments, becoming more and more ponderous and controversial."
"According to this model, the universe brims with energy that acts like a cosmic molasses, imbuing the particles that move through it with mass, the way a bill moving through Congress attracts riders and amendments, becoming more and more ponderous and controversial."
Umm, sure, that's a sensible analogy! :)
Anyway, that got me thinking of some more analogies with a political/ideological bent:
The Higgs field acts like a cosmic molasses, imbuing particles that move through it with mass...
...like a wolf traipsing carefree through the forest, only to get its paws progressively caught up in traps set illegally by hunters.
...like a gun that moves from dealer to dealer, impeded by endless background checks.
...like a all natural tomato growing for generations only to get progressively genetically modified by vicious genetic engineers.
...like a happy healthy child driven to autism by an extensive vaccine regimen.
...like a investment banker hampered by bureaucratic regulations.
...like a factory hit with repeated union strikes.
...like a group of workers progressively worked into the ground by management.
And some other non-political/ideological ones:
...like a PI who receives several e-mails a minute about how the latest seminar series has been canceled.
...like an unadorned Cheeto that picks up that cheesy powder on its trip through the Cheeto factory.
...like an adult working her or his way through life, only to be slowed by children, mortgage payments, and a variety of home maintenance projects.
Sunday, October 6, 2013
Impact factor and reading papers
I just read this paper on impact factor, and it makes the point that impact factor of a journal is a highly imperfect measure of scientific merit, but that the reason that the issue persists is that hiring committees fetishize publications in high impact journals. The authors say that this results in us scientists ceding our responsibilities for gauging scientific merit largely to a handful of professional editors, most of whom have very little experience as practicing scientists. All true. The real question is: why does this happen? Surely we as scientists would not have let things get so bad, especially when hiring decisions are so very important, right?
I think the issue comes down to the same basic issue that is straining the entire scientific system: the expansion of science and the accompanying huge proliferation in the amount of scientific information. Its a largely positive development, but it has left us straining to keep up with all the information that's out there. I think it's likely that the majority of faculty on hiring committees are just too busy to read many (if any) of the papers of the candidates they are evaluating, often because there's just not enough time in the day (and probably because most papers are so poorly written). So they rely on impact factor as some sort of gauge of whether the work has "impact". So far, pretty standard complaint. And the solutions are always the same boring standard platitudes like "We must take action and evaluate our colleagues work on its merits and not judge the book by its cover, blah blah blah." People have been saying this for years, and I think the reason this argument hasn't gone anywhere is that while it sounds like the right thing to say, it doesn't address the underlying problem in a realistic way.
For another way of looking at the issue, put yourself in the same position as this member of the hiring committee. What would you do if you had to read hundreds of papers, many highly technical but also not in your field of expertise, while running an entire lab and teaching and whatever else you're supposed to do? You would probably also look for some other, more efficient way to gauge the relative importance of these papers without actually reading them. That's why the research seminar is so useful, because it allows you to get a quick sense of what a scientist is all about. Which raises the question that I often have, which is why do we bother writing all these papers that nobody reads? My personal feeling is that we have to reevaluate how we communicate science, and that the paper format is just not suitable for the volume of information we are faced with these days. In our lab, we have slidecasts, which are 10 minute or less online presentations that I think are at least a step in the right direction. But I have some other ideas to extend that concept further. Anyway, whatever the solution, it needs to get here soon, because I think our ability to be responsible scientists will depend on it.
I think the issue comes down to the same basic issue that is straining the entire scientific system: the expansion of science and the accompanying huge proliferation in the amount of scientific information. Its a largely positive development, but it has left us straining to keep up with all the information that's out there. I think it's likely that the majority of faculty on hiring committees are just too busy to read many (if any) of the papers of the candidates they are evaluating, often because there's just not enough time in the day (and probably because most papers are so poorly written). So they rely on impact factor as some sort of gauge of whether the work has "impact". So far, pretty standard complaint. And the solutions are always the same boring standard platitudes like "We must take action and evaluate our colleagues work on its merits and not judge the book by its cover, blah blah blah." People have been saying this for years, and I think the reason this argument hasn't gone anywhere is that while it sounds like the right thing to say, it doesn't address the underlying problem in a realistic way.
For another way of looking at the issue, put yourself in the same position as this member of the hiring committee. What would you do if you had to read hundreds of papers, many highly technical but also not in your field of expertise, while running an entire lab and teaching and whatever else you're supposed to do? You would probably also look for some other, more efficient way to gauge the relative importance of these papers without actually reading them. That's why the research seminar is so useful, because it allows you to get a quick sense of what a scientist is all about. Which raises the question that I often have, which is why do we bother writing all these papers that nobody reads? My personal feeling is that we have to reevaluate how we communicate science, and that the paper format is just not suitable for the volume of information we are faced with these days. In our lab, we have slidecasts, which are 10 minute or less online presentations that I think are at least a step in the right direction. But I have some other ideas to extend that concept further. Anyway, whatever the solution, it needs to get here soon, because I think our ability to be responsible scientists will depend on it.
Saturday, October 5, 2013
Why are figures hard to read?
In a previous post, I talked about how figure legends are quite useless. But that got me thinking about figures more generally. Gautham and I have both been talking about how sometimes we read papers first without even looking at the figures. I like this style of reading because it allows you to pay attention to the authors' arguments without interruption. Of course, the problem is that you don't actually see the data they're referring to. But if you trust that their interpretation of their graph or picture or whatever is correct (otherwise they wouldn't be showing it, right?), then there's really no point in looking at the figure, right? Hmm...
So why am I avoiding figures? I think the reason is that there are a lot of cognitive hurdles to doing so. Many papers will have some sort of an interpretative statement, like this "Knocking down the blah-1 gene led to a substantial increase in cells walking to the side of the plate and turning purple (Fig. 2D)." Okay, but then you look at the figure, and it will probably have some sort of bar graphs of knockdown efficiency and some cell-walking-turning-purple assay readout. Then I have to think about what those charts mean and how they did the assay and whether that makes sense and... you get the idea. It requires a lot of thinking, and by the time you understand it, I defy anyone to go right back to the main text and regain their previous train of thought.
What to do? Well, this seems to be far less of a problem when you're giving a talk. Why? I think it's because you have the opportunity to spend time with the figure and explicitly show people what you want them to get out of it as an integrated part of your narrative. In a paper, I think one of the things we could do is to adopt things like sparklines (ala Tufte), which are inline graphics. Here's an example lifted straight out of Wikipedia: The Dow Jones index for February 7, 2006  . Let's take our previous example. What if I said something like "Knocking down the blah-1 gene
. Let's take our previous example. What if I said something like "Knocking down the blah-1 gene  led to a substantial increase in cells walking to the side of the plate and turning purple
led to a substantial increase in cells walking to the side of the plate and turning purple  . Simple and informative, and inline, so there's no cognitive interruption. (Note: they came out bigger than I wanted, but it's hard to deal with this pesky Blogger interface.) Does this have all the information that a figure would have? No. But it conveys the main point. And probably with slight larger graphics, you could include a lot of that information. As it is with most figures, there's so much missing information anyway in terms of how one actually made the figure that it's not like it's all that complete to begin with. The best way to deal with that is if we adopted a writing style in which we didn't just give an interpretation, but actually took the time to explain what we did. That's hard within the writing limitations imposed by the journals, though.
. Simple and informative, and inline, so there's no cognitive interruption. (Note: they came out bigger than I wanted, but it's hard to deal with this pesky Blogger interface.) Does this have all the information that a figure would have? No. But it conveys the main point. And probably with slight larger graphics, you could include a lot of that information. As it is with most figures, there's so much missing information anyway in terms of how one actually made the figure that it's not like it's all that complete to begin with. The best way to deal with that is if we adopted a writing style in which we didn't just give an interpretation, but actually took the time to explain what we did. That's hard within the writing limitations imposed by the journals, though.
 . Simple and informative, and inline, so there's no cognitive interruption. (Note: they came out bigger than I wanted, but it's hard to deal with this pesky Blogger interface.) Does this have all the information that a figure would have? No. But it conveys the main point. And probably with slight larger graphics, you could include a lot of that information. As it is with most figures, there's so much missing information anyway in terms of how one actually made the figure that it's not like it's all that complete to begin with. The best way to deal with that is if we adopted a writing style in which we didn't just give an interpretation, but actually took the time to explain what we did. That's hard within the writing limitations imposed by the journals, though.
. Simple and informative, and inline, so there's no cognitive interruption. (Note: they came out bigger than I wanted, but it's hard to deal with this pesky Blogger interface.) Does this have all the information that a figure would have? No. But it conveys the main point. And probably with slight larger graphics, you could include a lot of that information. As it is with most figures, there's so much missing information anyway in terms of how one actually made the figure that it's not like it's all that complete to begin with. The best way to deal with that is if we adopted a writing style in which we didn't just give an interpretation, but actually took the time to explain what we did. That's hard within the writing limitations imposed by the journals, though.Sunday, September 29, 2013
PubReader format rocks!
By the way, is it just me, or does the new PubReader format on NIH PubMedCentral totally kick butt? I can't help but wonder why journals have not gone to something like that. The best thing is that you can quickly glance at a figure–any figure–rapidly by just hovering quickly over the figure thumbnails at the bottom of the page. You read, see a reference to a figure, and can quickly take a peek (and then quickly come back). Why it took somebody 15 years to do something like this is anyone's guess, but I sure am grateful somebody finally did it.
Oh, and the best part is that it's open access! These are the free versions of the articles you have to submit to PubMedCentral.
Oh, and the best part is that it's open access! These are the free versions of the articles you have to submit to PubMedCentral.
What's the point of figure legends?
Figures in journals have figure legends. But what's the point of these figure legends? In my mind, there is little point. At best, they can serve as a sort of "materials and methods" for the figure. But if people have to read something to actually make sense out of what's in the figure, then you've already lost the battle.
This happens all the time. Take a look at this figure.
Now in part C, there are some black data points and some red data points. What's the difference? What are the authors trying to tell you? Well, here's what the text is trying to tell me I should get from this figure:
To which the paper replies:
Here's another more biological example:
The paper tells you that
On a more general note, I think that sometimes authors approach their paper as "work" for the reader, that it's somehow good to struggle through a paper to understand it's meaning. Many biology papers with endless combinatorial gels have this quality, like one of those "If Harry can't sit next to Sally on Thursday but wants to sit next to Alice on Wednesday" kind of problems. I think that the notion that this is somehow "good medicine" is ridiculous. A scientific paper's goal is not to be the crossword in the Sunday Times. It's a vehicle for communicating results as clearly and quickly as possible. Our goal as authors is to achieve that result as best possible. It's not an easy task, but that's our job.
This happens all the time. Take a look at this figure.
Now in part C, there are some black data points and some red data points. What's the difference? What are the authors trying to tell you? Well, here's what the text is trying to tell me I should get from this figure:
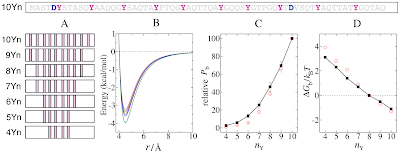
As for other EAD sequences studied before, transactivation rises in a nonlinear manner with nY (Fig. 1C, red circles), demonstrating that multiple Ys act together in a cooperative manner.Okay, so the text is telling us to look at this figure to see some sort of non-linear phenomenon. Okay, but presumably these black things mean something also, right? Some sort of comparison? Wait, the legend must know!
To which the paper replies:
(C) Effect of Y number nY on transactivation and simulated binding. Relative transcriptional activity of the EAD peptides (open red circles) was determined under sub-saturating conditions (Methods and Text S1) relative to 10Yn activity (arbitrarily set to 100). Red error bars for the experimental data indicate SEM. The relative Pb(nY) values (filled black squares) are normalized by the Pb for 10Yn [nY = 10, actual simulated (absolute) Pb(10) = 0.43]. The black error bars mark standard deviations among ten independent simulations.Ouch. This highlights the problem with this format: the paper is telling me some relatively high level interpretation, then I look to the figure, but I can't make sense of it, so then I have to look at the legend, which is obfuscated to the point of utter incomprehensibility. Why would you make your reader work so hard?
Here's another more biological example:
The paper tells you that
Ectopic COT expression in A375 and SKMEL28 cells also conferred decreased sensitivity to the MEK inhibitors CI-1040 and AZD6244, suggesting that COT expression alone was sufficient to induce this phenotype (Fig. 4c, 4d, Supplementary Fig. 17).Hmm. Something about Ectopic COT expression. But I don't see any explanation of MEK1, or MEK1 DD (turns out they are negative and positive controls). In fact, just looking at this figure, I would have no idea that this was about overexpression whatsoever. Nor would I be able to tell what's going on with SKMEL28, which isn't even on here (probably in Supp. Fig. 17). And the legend gives us:
ERK phosphorylation in A375 expressing indicated ORFs following treatment with DMSO or 1µM of PLX4720, RAJ265, CI-1040 or AZD6244.Again, ouch! What are all of these different drugs doing? Which is the control (DMSO)? Which one inhibits what? What should I be drawing from this figure? This figure is a bit better than my other example, because the audience of the paper might actually have some notion of what this is all about due to common background knowledge, but the cognitive demands for understanding the point of this experiment for both the initiated and uninitiated are unreasonably high. Just imagine what a few simple annotations or diagrams could do here...
On a more general note, I think that sometimes authors approach their paper as "work" for the reader, that it's somehow good to struggle through a paper to understand it's meaning. Many biology papers with endless combinatorial gels have this quality, like one of those "If Harry can't sit next to Sally on Thursday but wants to sit next to Alice on Wednesday" kind of problems. I think that the notion that this is somehow "good medicine" is ridiculous. A scientific paper's goal is not to be the crossword in the Sunday Times. It's a vehicle for communicating results as clearly and quickly as possible. Our goal as authors is to achieve that result as best possible. It's not an easy task, but that's our job.
Monday, September 16, 2013
Images in presentations
Since Gautham is on the subject of presentation pet peeves, I thought I'd bring up one of my own (along with some solutions).
We do a lot of imaging in the lab, and one of the best things about it is that it can produce compelling images that are fun to show off at a talk. Nothing like a few oohs and aahs to bring the audience back from the dead...
But one thing that always bugs me in talks is the dreaded "Well, you can't really see it so well here, but trust me, it looks really good on my screen..." (and yes, it's happened to me many times). This is often accompanied by a valiant but typically doomed attempt to show the image on the laptop to someone in the audience to try and corroborate the claim. What are the common reasons that images look bad on the screen? Here are a couple along with some suggestions.
1. Image features are too small to see. We have a lot of RNA FISH images that we like to show off in talks. Which of course brings up a problem: the RNA spots are really small, and often we want to show off multiple cells. Or, even worse, comparisons between fields of cells. The temptation is to put those fields next to each other and shrink down the image. The problem then is that the images become so compressed that nobody can see the spots anyway. So then what's the point? If you have an image scale issue, one solution is to break the problem down. Show the outermost scale to get people oriented (along with markers that are actually visible at that scale), then show a little box around the area, and then zoom in.
2. Merges. Gautham covered this nicely. Merges usually just look really bad. Try not to use them. There are a few situations in which they can be useful, but they are rare.
3. Contrast is to low. This is actually an issue that's even more pronounced in print. The problem is that stuff that's obvious when you're sitting right next to your screen is hard to see on the projector. I think this is because when you are close up and have time to pay attention, it's easier to focus on what are actually rather subtle features in the image.
4. Settings on Apple Laptops
Here's another seemingly arcane but often crucial little trick. Despite my best efforts to adhere to the above rules, a few years ago I noticed that after I updated my laptop's OS, I found myself repeatedly saying "Well, somehow this isn't looking so good on the screen, but what you should see is..." Very embarrassing! And I just could not for the life of me figure out why my images were looking so crappy on the screen. Then, after some digging, I figured it out. Turns out that when you plug in a external projector (or monitor or whatever) to your Mac, it chooses a color profile for that device. This governs how the colors look on the screen. The issue is that a couple years back, Apple updated things so that the default color profile when you connect has terrible contrast for many images. The solution is to open "Displays" in System Preferences, then go over to the output screen, click on the "Color" tab, then uncheck the box marked "Show profiles for this display only", and then select "sRGB IEC61966-2.1". Anyway, after I did that, all those weird problems went away. But only after dealing with the first 3 items...
Time
I was in a shower at a hotel this morning, and I was wondering whether it was worth the time to put the cap back on the little bottle of shampoo after I used it, given that whoever cleaned the room afterwards would most likely just throw it away anyway. What if you added up all those little microseconds of wasted time in your life? It would probably add up to about as much time as it took to write this blog post... :)
Wednesday, September 11, 2013
A case against laser pointers for scientific presentation
- Gautham
Part 2 of a three part blog post. First part was on color merges. Perhaps there will be a third on 3d plots, which is the least pressing because it is the least common and the least damaging.
Laser pointers are the most common, and they invariably make for a poorer presentation than without it. Here are the three fundamental problems with the use of laser pointers in talks.
So what are good pointing devices to use?
Part 2 of a three part blog post. First part was on color merges. Perhaps there will be a third on 3d plots, which is the least pressing because it is the least common and the least damaging.
Laser pointers are the most common, and they invariably make for a poorer presentation than without it. Here are the three fundamental problems with the use of laser pointers in talks.
- Laser pointers do not point at things, they point on things. In fact, the term "laser pointer" is a misnomer. It is just a laser point. It does not produce a pointer, such as an arrow or a pointing hand. To highlight a feature with the laser you must overlap it with your extremely bright laser point, completely obscuring the feature as well as readjusting your eye's dynamic range to make the actual features on the slide harder to see. Laser pointer companies pride themselves in outdoing each other with brightness, and audiences are impressed by green laser pointers, the most distracting of all. This is missing the ... "point". When attempting to highlight punctate features the problem finds its ultimate aggravation (and don't get me started when laser pointing on color-merged puncta. Save our souls!). When a laser pointer is activated, its dot is invariably the brightest thing for your eye to look at. In summary, if you use a laser pointer, we in the audience see the laser dot. Not the thing you want us to see.
- The laser pointer is the only moving thing in our field of view. Suppose you were looking at a completely stationary picture and something moves in it. We are going to pay attention to the part that moves, not the part that sits still. Otherwise we have to use extreme concentration to suppress our instinct to focus on the stationary medium-intensity background instead of the in-motion brightly colored laser spot. In doing so, we are attempting to suppress an evolved instinct that saved our ancestors' necks countless times. This is another reason why when a presenter uses a laser pointer, we focus on the laser point, not whatever it is he/she is trying to point at.
- A laser pointer fosters the detachment of the presenter from the presentation. This is unfortunately the standard presentation mode by the entire community without anyone ever having made a conscious choice about it and has broader sources than just laser pointers. The modern scientific talk has turned into a kind of slide-cast where we all look at the slides and listen to the disembodied voice of the presenter. The two are sporadically connected by a device that projects an unseen beam ending in a bright dot. When a presenter wants to point at a feature, he/she will often step back from the presentation, further increasing the detachment. Is that how you want to learn? From a slide or from a person? Would you be happy if that's how your kids got taught in school? Do you want a talk where the slides are an aid and extension of the presenter, or one in which the presenter's role is to provide commentary on slides? I decided many years ago that scientists are more interesting that powerpoints, so if a presenter forces me to pick between looking at the slides and looking at them (which most do), I always look at them and mostly ignore whatever is projected.
So what are good pointing devices to use?
- Your hand. Best pointing device invented. Unlike the laser pointer, you can point at stuff with this, not just on stuff. Walk into your slide if necessary. It is your helper, not the other way around. Stay connected to it.
- A stick. Another classic pointing device. It extends your hand nicely to get to some of those plots you put at the top of your slide.
- Arrows in the presentation. If you can't do the first two, which may be if you have to talk in an enormous lecture hall and your podium has been unwisely (but commonly) placed far from the projection screens, you can animate an arrow to appear on your slide when you want to call attention to something. This requires you to think beforehand about what you want to call attention to.
What is a good use of a laser pointer?
Turns out they make passable lasers for optical applications. You can use them to align your complicated optical experiment, or even do fluorescence microscopy (I've used them for both). It is also apparently the case that the laser dot is very entertaining to cats and they chase after it like it is prey, which coming to think of it is a lot like what we humans do (points 1 and 2 above), so they sell these pointers at pet stores. Unlike humans, though, my brother reports that his cat eventually tired of the novelty.
Monday, September 9, 2013
A case against color-merges to show colocalization
- Gautham
Seeing as a recent post has revealed to the public my distaste for certain common scientific data display and presentation techniques (laser pointers, 3d plots and color-merges), it seemed a good time to explain why I'm not into these things. The color-merge comes first. I'll explain why I think it is a too-clever trick that is best replaced with showing each image separately.
It is often intended to show that signals in two different imaging channels, usually two fluorescent dyes or proteins, arise from the same underlying objects, or "colocalize." Scientists commonly find it necessary to take the two images, invariably collected in black and white, false-color them and superimpose them using some form of color-combining (usually an additive color model). Then you show a figure where you say that A and B where false colored red and green and look, in the combination all the features are purple. Or is it orange? Actually, what is the sum of red and green? Or are we subtracting red and blue from white? Isn't red plus green black?
I think this is the root of the problem with color merging for co-localization. We have no innate skill at combining colors. I have to look up a color wheel, or remember my elementary school painting class, to determine what the combination of two colors is supposed to be. Take a look at the wikipedia page for http://en.wikipedia.org/wiki/Cmyk and ask yourself if you get any idea for the full color print by looking at the CMYK decomposition. Aside from painting, when do we ever combine colors? When did our ancestors' lives or reproductive potential depend on their ability to sum red and green?
On the other hand, we are good at recognizing visualizing patterns. We do so innately and struggle to program a computer to match our capabilities. We can recognize a person's face in a caricature or in a minimalist hand drawing or despite considerable digital distortion and no matter what else is in the background. When two images are images of the same thing, we can tell. Plus, there is no need for chromatic-shift correction to make the point.
A corollary of these arguments is that color-merging is actually not too bad at showing lack of colocalization or simply to overlay two totally different types of images. For example, if you show diffuse immunofluorescence in one color and punctate RNA-FISH in another color and add them up. This does not demand that we add colors, and is a useful help for us to judge the spatial relationships between these two stains. It is analogous to having a foreground and a background in a picture. You can also color-merge two different punctate signals that do not co-localize and not do much harm (though it may be a wasted effort).
In short, if it can be avoided, don't require us to add colors.
Here are some other things that are sacrificed when color merging:
Seeing as a recent post has revealed to the public my distaste for certain common scientific data display and presentation techniques (laser pointers, 3d plots and color-merges), it seemed a good time to explain why I'm not into these things. The color-merge comes first. I'll explain why I think it is a too-clever trick that is best replaced with showing each image separately.
It is often intended to show that signals in two different imaging channels, usually two fluorescent dyes or proteins, arise from the same underlying objects, or "colocalize." Scientists commonly find it necessary to take the two images, invariably collected in black and white, false-color them and superimpose them using some form of color-combining (usually an additive color model). Then you show a figure where you say that A and B where false colored red and green and look, in the combination all the features are purple. Or is it orange? Actually, what is the sum of red and green? Or are we subtracting red and blue from white? Isn't red plus green black?
I think this is the root of the problem with color merging for co-localization. We have no innate skill at combining colors. I have to look up a color wheel, or remember my elementary school painting class, to determine what the combination of two colors is supposed to be. Take a look at the wikipedia page for http://en.wikipedia.org/wiki/Cmyk and ask yourself if you get any idea for the full color print by looking at the CMYK decomposition. Aside from painting, when do we ever combine colors? When did our ancestors' lives or reproductive potential depend on their ability to sum red and green?
On the other hand, we are good at recognizing visualizing patterns. We do so innately and struggle to program a computer to match our capabilities. We can recognize a person's face in a caricature or in a minimalist hand drawing or despite considerable digital distortion and no matter what else is in the background. When two images are images of the same thing, we can tell. Plus, there is no need for chromatic-shift correction to make the point.
A corollary of these arguments is that color-merging is actually not too bad at showing lack of colocalization or simply to overlay two totally different types of images. For example, if you show diffuse immunofluorescence in one color and punctate RNA-FISH in another color and add them up. This does not demand that we add colors, and is a useful help for us to judge the spatial relationships between these two stains. It is analogous to having a foreground and a background in a picture. You can also color-merge two different punctate signals that do not co-localize and not do much harm (though it may be a wasted effort).
In short, if it can be avoided, don't require us to add colors.
Here are some other things that are sacrificed when color merging:
- For a co-localization merge, it is difficult to deal with the background and the signal in each image in a consistent way and still deliver your message. Unless the two images are linear transformations of each other, it will be impossible for the whole co-localized image to be the same color, despite perfect co-localization of the signal.
- Co-localization merge requires that not only the positions of the features be the same but that the ratio of intensities of the features in the two channels be always the same. It is quite difficult to know if a little bit of red has been added to green.
- You give up the freedom to independently contrast and colormap your separate images, which is very important when dealing with data that contains features at widely different intensities. You can then use color very effectively to encode intensity within each channel's image, and let our ample pattern recognition capabilities judge the overlap between channels.
Sunday, September 8, 2013
Courage in education
I just read an interesting article on NYTimes.com about supporting and encouraging women at Harvard Business School through a series of somewhat radical changes. These changes included what some of the naysayers called "social engineering" in order to create an environment more friendly to women (these mostly sounded good things to me). In the article, they profiled a woman who ended up giving the class's graduation speech. The theme of the speech was courage, and how much courage it takes to change things that are wrong. She particularly mentioned the courage of the one administrator, Ms. Frei, who took on the culture at Harvard Business School and decided to change it. Having read the article and considering the history and powerful interests I'm sure are at play at a place like Harvard Business School, I think that calling that move courageous is pretty accurate, and I think Ms. Frei's example is pretty inspiring. Another thing I took from the article is that real change usually involves pissing some people off. I'm wondering if maybe the trick is to know how to not to piss off just enough people that your ideas actually make it to reality.
Which led me to wonder where are areas in my job where I could be more courageous? I feel like in research, being courageous is sort of par for the course, since the whole point of any research project is to learn something that hopefully changes the way people think. (I think there are some places where a little courage could change how we communicate science... maybe in another post...) What about education, though? I think this is an area that is ripe for people being more courageous. I think that many people realize that the way we teach probably needs a fundamental overhaul–bottom line is that most students just aren't learning as much as they can or should. Doesn't it seem odd to you that we teach largely in the same way we have for the last several hundred years? Do we do anything the same way we did hundreds of years ago?
But change of the scale that at least I think we should engender will definitely piss off a lot of people, and will require someone courageous at the upper levels of university administration to implement/force down people's throats. You could imagine these sorts of things growing from the bottom up–i.e., from the faculty–but that has two problems: 1. even faculty who believe in this (like me) have so many demands on their time that we tend to largely revert to the default way of teaching; and 2. given that education is an endeavor that now typically occurs across departments, some sort of higher level coordination is needed.
Then again, maybe we don't have to wait that long. In research, especially in biology, I feel like changes in technology can force changes even upon those unwilling to accept them. Online education is perhaps just such a disruptive force. It's early, but I think it could be a chance to fundamentally reshape higher education. And it definitely pisses a lot of people off, which is a good sign... :)
Which led me to wonder where are areas in my job where I could be more courageous? I feel like in research, being courageous is sort of par for the course, since the whole point of any research project is to learn something that hopefully changes the way people think. (I think there are some places where a little courage could change how we communicate science... maybe in another post...) What about education, though? I think this is an area that is ripe for people being more courageous. I think that many people realize that the way we teach probably needs a fundamental overhaul–bottom line is that most students just aren't learning as much as they can or should. Doesn't it seem odd to you that we teach largely in the same way we have for the last several hundred years? Do we do anything the same way we did hundreds of years ago?
But change of the scale that at least I think we should engender will definitely piss off a lot of people, and will require someone courageous at the upper levels of university administration to implement/force down people's throats. You could imagine these sorts of things growing from the bottom up–i.e., from the faculty–but that has two problems: 1. even faculty who believe in this (like me) have so many demands on their time that we tend to largely revert to the default way of teaching; and 2. given that education is an endeavor that now typically occurs across departments, some sort of higher level coordination is needed.
Then again, maybe we don't have to wait that long. In research, especially in biology, I feel like changes in technology can force changes even upon those unwilling to accept them. Online education is perhaps just such a disruptive force. It's early, but I think it could be a chance to fundamentally reshape higher education. And it definitely pisses a lot of people off, which is a good sign... :)
Saturday, September 7, 2013
Rare Gautham sighting...
The sight of Gautham actually using a laser pointer was so bizarre that we had to record the event:
From group meeting yesterday. In other news, Gautham requested merged imaged channels and 3D plots. And reportedly said "Eh, statistical analysis, schmanalysis! Who needs that stuff, anyway." But you'll just have to take my word for it...
From group meeting yesterday. In other news, Gautham requested merged imaged channels and 3D plots. And reportedly said "Eh, statistical analysis, schmanalysis! Who needs that stuff, anyway." But you'll just have to take my word for it...
Saturday, August 31, 2013
GO analysis by "greedy" assignment of genes to categories
- Gautham
This post proposes a simple method to analyze gene hits from a screen in terms of GO categories, provides an example from the literature, and links to R code.
It is easy to find many very sophisticated algorithms and frameworks for analyzing the results of high throughput screens in the context of biological annotations to categories like the Gene Ontology (GO). The result is almost always p-values and enrichment scores for all GO categories, and it is common practice to then report the "most significant" (lowest p-value) terms. There are often a very large number of terms that meet reasonable significance criteria even after multiple hypothesis testing correction. This may be partly due to improvement in experimental methods, so that now even small effects result in statistically significant gene hits. Many comparative RNA-Seq experiments yield thousands of differential expression gene hits.
It is tough to get a feeling for the make-up of the hit gene list by a vanilla GO analysis because of the following problems:
- Screening by p-value tends to give large categories. This is for the same reason that screening differential expression by p-value tends to give genes with high average expression.
- Screening by enrichment gives very small categories. Again, for the same reason as screening differential expression by fold change tends to give genes with very low average expression.
- GO terms are not mutually exclusive, and hit lists tend to be made up of overlapping GO categories whose high rank on the list is due to a common set of hits.
The first two problems are common to any differential expression screen, but the overlap problem is unique to category analysis. Without exaggeration, these are the top 10 hits by p-value for a set of gene hits from a screen we did in the lab:
- multicellular organismal process
- anatomical structure morphogenesis
- system development
- multicellular organismal development
- anatomical structure development
- developmental process
- organ morphogenesis
- signaling
- skeletal system development
- nervous system development
More sophisticated GO analyses take into account the graph structure of GO to try to get around this problem, but it seemed like something very simple could do the trick instead. Here is a proposal:
- Decide the scale of GO terms you want to consider at the outset. Only consider GO terms that have more than some minimum number of genes annotated to it. The larger the cutoff, the broader the picture you will get.
- Find the GO term with the highest concentration of hits. This category has the greatest number of hits in the screen per annotated gene.
- Assign these hit genes to that GO category. Remove these genes from further consideration.
- Repeat 2,3,4 but considering only hits that remain unassigned. Continue until all hits are assigned, or you have as many GO terms as you want. Usually, the parent biological_process category will eventually get selected and sweep up all remaining genes.
Step 1 forces you to explicitly choose your resolution. Step 2 seeks to find categories that are as likely as possible to be actually involved in the biology you are studying. Step 3 bravely (or foolishly) assigns genes as doing the most enriched function they are annotated for, and Step 4 eliminates the GO overlap problem. This kind of algorithm is commonly known as "greedy," so we refer to it sometimes as greedy GO.
To illustrate we used the data set generated by Marks et al. 2012 (PMID: 22541430). These authors looked at the transcriptome differences between mouse ES cells maintained in serum-containing media or media containing two small molecule signaling inhibitors. They found nearly 1500 genes significantly more highly expressed in 2i media than in serum, and present their own category analysis (PANTHER database) in their Fig. 1c. Using the BioConductor-maintained org.Mm.eg.db annotation database for mouse as a source for GO annotations, here is a breakdown by the greedy method of the "high in 2i" gene hits from Marks et al. Cell 2012:
| Num.hits.assigned | Num.hits | Num.genes | |
|---|---|---|---|
| cellular lipid metabolic process | 73 | 73 | 495 |
| regulation of immune system process | 49 | 54 | 393 |
| cellular nitrogen compound biosynthetic process | 40 | 44 | 328 |
| ion transmembrane transport | 41 | 45 | 374 |
| carbohydrate metabolic process | 43 | 58 | 414 |
| neuron projection development | 33 | 38 | 382 |
| biological_process | 1004 | 1283 | 12243 |
These are not the top hits, or handpicked in any way. They are the complete output of our greedy algorithm. No p-value threshold has been set here. In fact, the method does not even compute p-values. Compare to Fig. 1c in the paper, and you'll see the categories mostly agree, and that we avoided duplicate entries for lipid metabolism. The approach can be augmented with p-values by running something like goseq, which calculates the p-values for every GO category.
The R code for the functions that do the computations is here:
https://gist.github.com/gauthamnair/6400111
A script that does an analysis of the Marks et al. data is below, showing a simple way to get GO annotations into the right format, by exploiting a function in the goseq package.
https://gist.github.com/gauthamnair/6400293
Subscribe to:
Posts (Atom)